Software
SpaceTracer
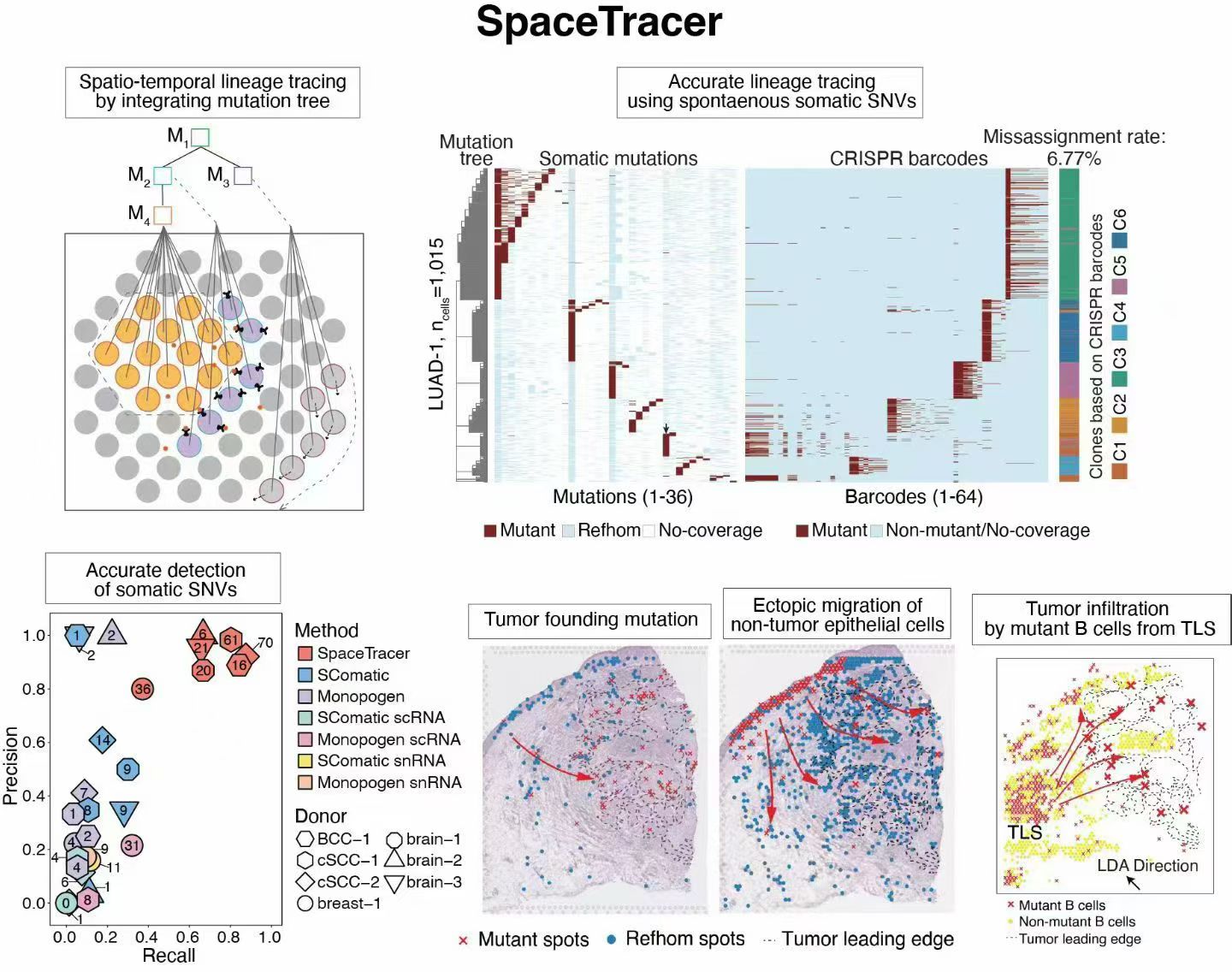
SpaceTracer is a computational framework that detects somatic single-nucleotide variants (SNVs) directly from spatial transcriptomics data and reconstructs cellular phylogenies in situ. By leveraging naturally occurring somatic mutations, it enables perturbation-free spatiotemporal lineage tracing and supports analyses of lineage spread, migration, and lineage-associated transcriptional and spatial dynamics in complex tissues.
PhyloSOLID
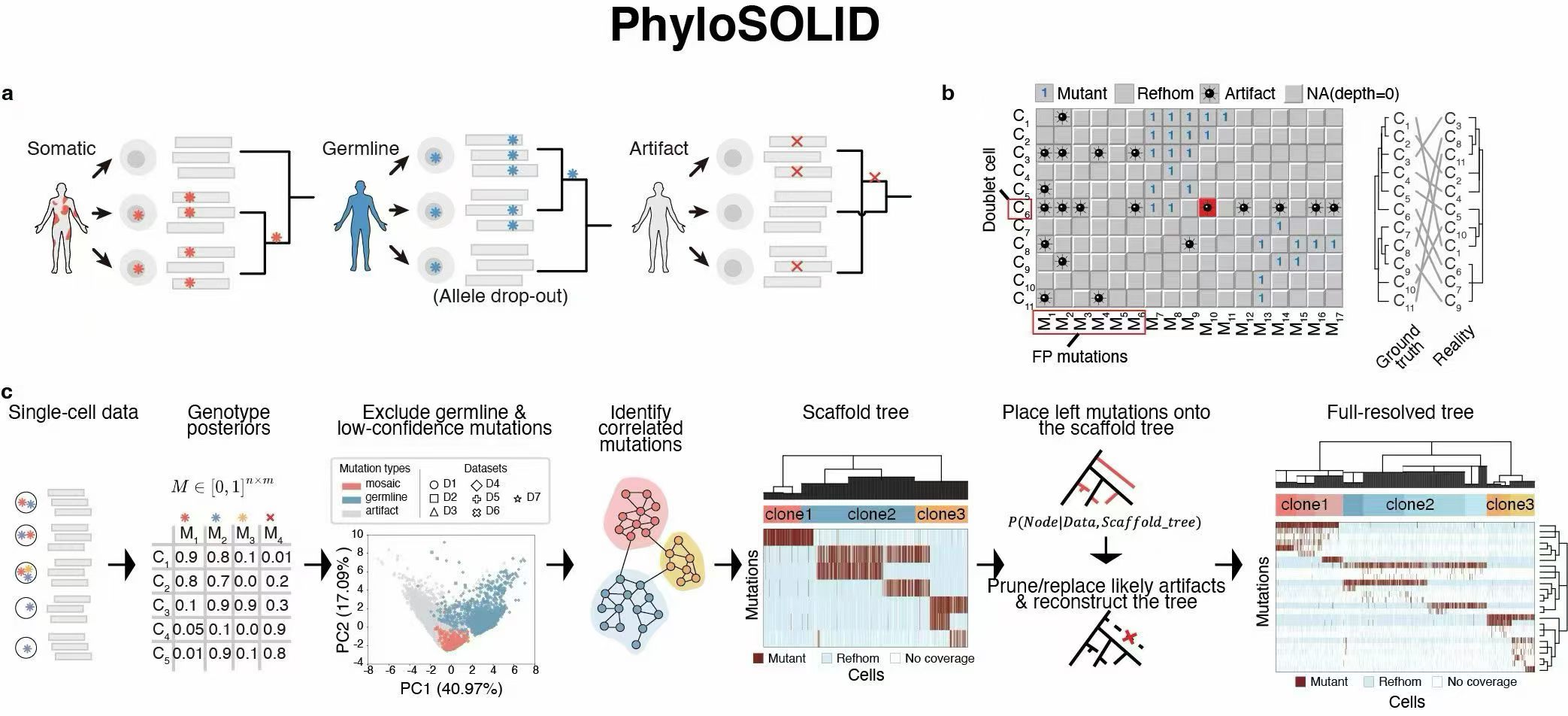
PhyloSOLID is a phylogenetic reconstruction algorithm designed for robust lineage inference from single-cell sequencing data despite high error rates and sparsity. It builds a high-confidence backbone tree from reliable mutations and progressively refines it using Bayesian modeling to reduce spurious phylogenetic signals and improve tree topology accuracy across scRNA-seq and scDNA-seq datasets.
BayesMonSTR
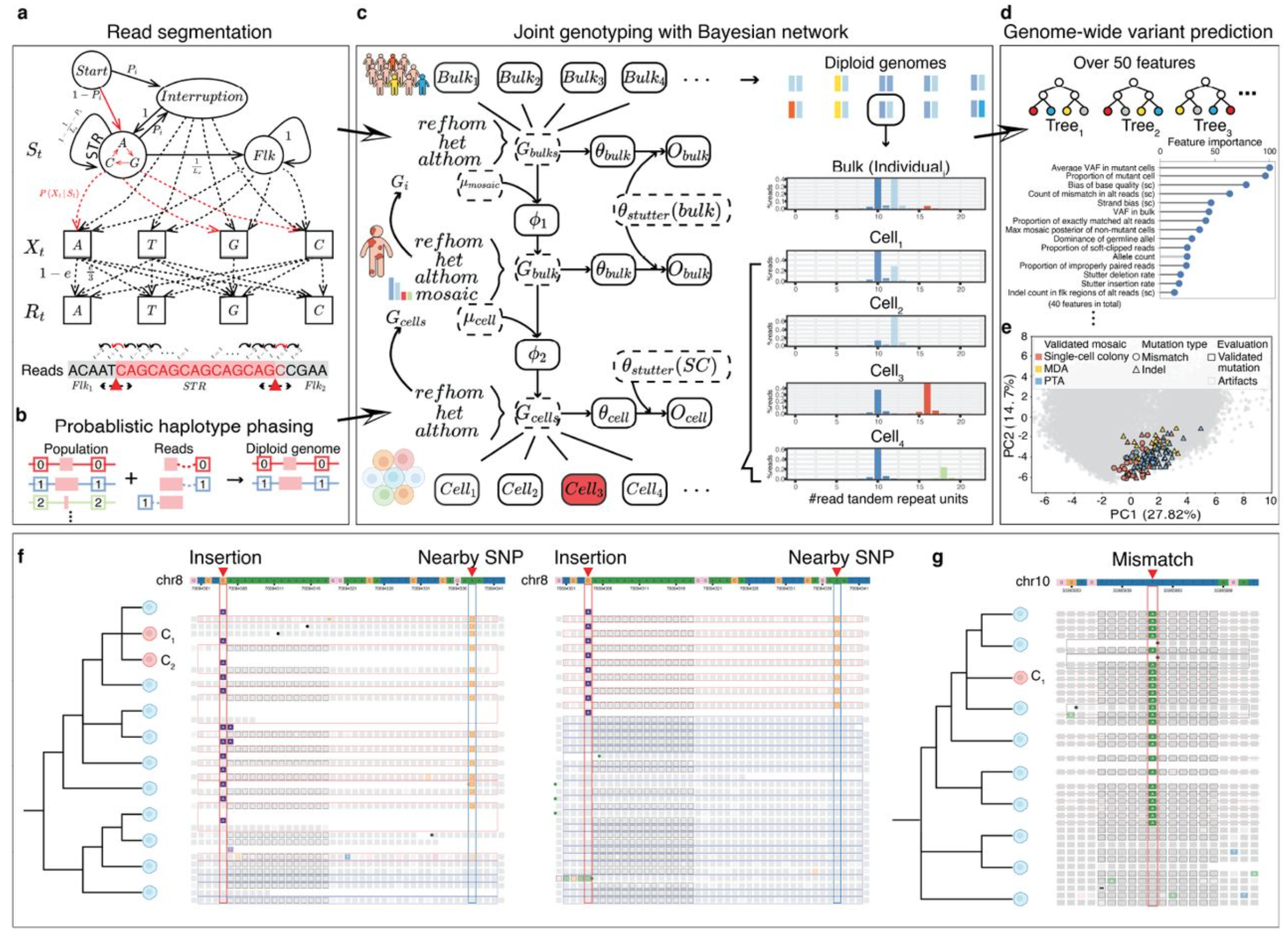
BayesMonSTR is a computational method for genome-wide detection of mosaic microsatellite (STR) mutations at single-cell resolution. It enables robust identification of mosaic STR insertions and deletions and facilitates downstream analyses of mutation burden and regulatory enrichment (e.g., transcription start sites and active enhancers) across tissues and aging.
MosaicForecast
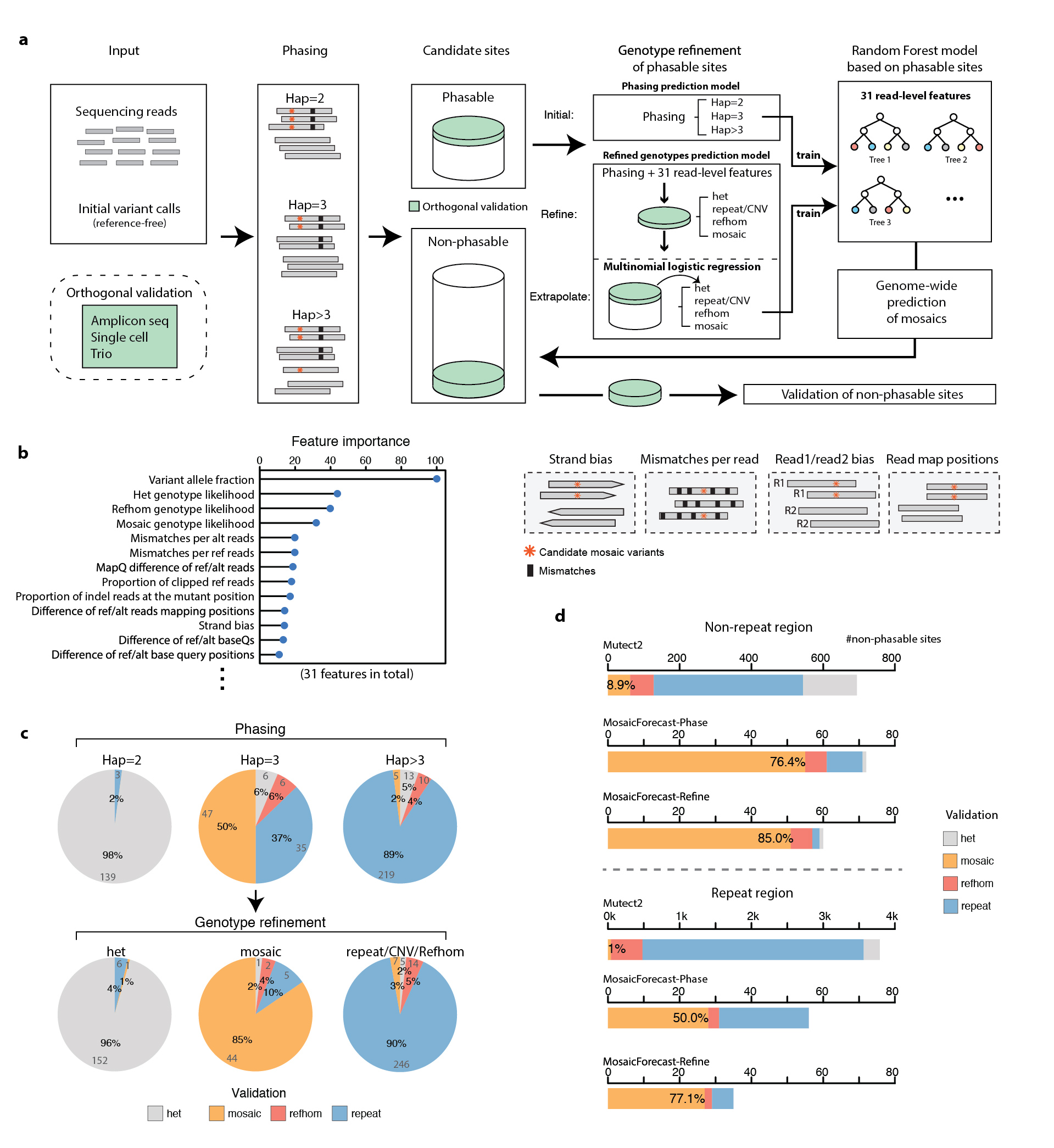
MosaicForecast was developed to detect mosaic point mutations and short indels from bulk sequencing data. It is a machine-learning method that leverages read-based phasing and read-level features to accurately detect mosaic single-nucleotide variants and indels, achieving a multifold increase in specificity compared with existing algorithms.
MosaicHunter
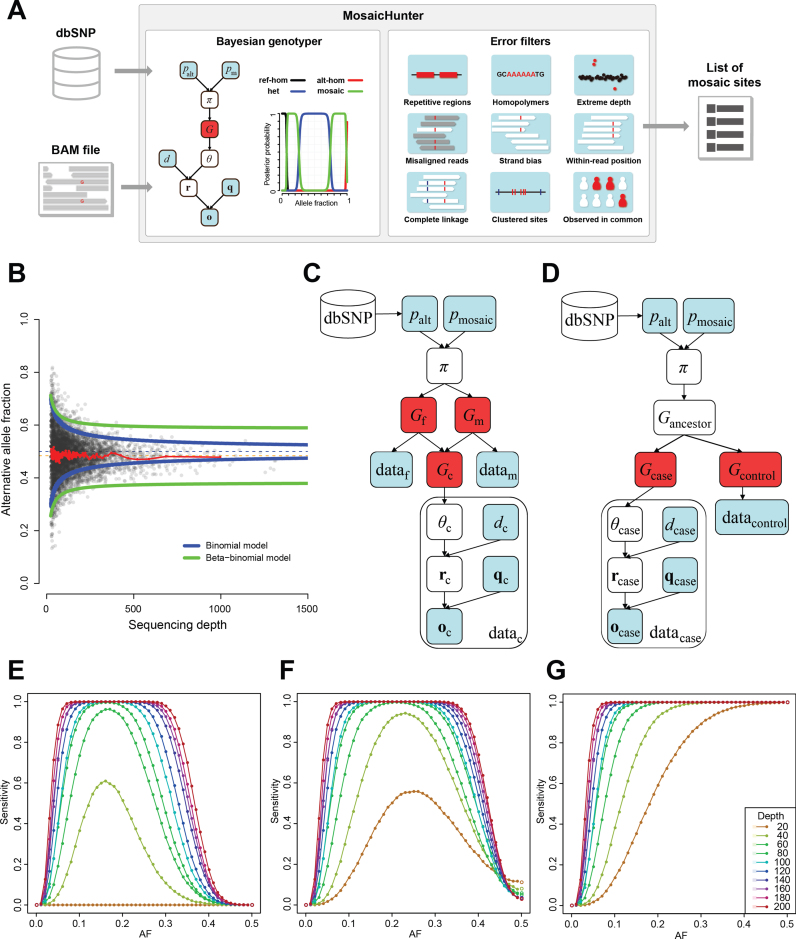
MosaicHunter is a bioinformatics tool that can identify SNMs in whole-genome and whole-exome sequencing data of unpaired samples without matched controls using Bayesian genotypers.