Outputs¶
This page summarizes important files generated by the current SpaceTracer workflow.
Key intermediate outputs¶
-
output_dir/cluster/cluster.txtandoutput_dir/cell_num.txt(or your configured externalcell_numfile) Cluster assignments and cell-number support for downstream genotype modeling. If you provide these files in config or skip early steps, these outputs may not be regenerated underoutput_dir. -
output_dir/bam_processing/IN.bam,output_dir/bam_processing/IN_filter.bamIn-tissue BAM and filtered BAM used bympileupand read-level feature extraction. -
output_dir/mpileup/raw_mpileup.txt,output_dir/mpileup/filter_mpileup.txt,output_dir/mpileup/umi_combine_chunk_manifest.tsvRaw/filter mpileup evidence and chunk manifest used byumi_combine. -
output_dir/umi_combine/spot.count.list.csv,output_dir/umi_combine/error.count.list.csvChunk-manifest files that point to per-chunk parquet outputs for spot/error evidence. -
output_dir/prior.txtPrior table used ingenotyping. -
output_dir/genotyping/genotyping_chunk_manifest.tsvPer-chunk genotype outputs (for exampleind_geno_filter.out,spot_geno.out,cluster_vaf.out) underoutput_dir/genotyping/<chunk>/. -
output_dir/spatial_feature/<sample>/spatial_feature_chunk_manifest.tsvSpatial feature chunk outputs. -
output_dir/read_feature/<sample>/read_feature_chunk_manifest.tsvRead-level feature chunk outputs. -
output_dir/mappability_feature/mappability_feature.txtMappability feature table. -
output_dir/RNA_feature/RNA_feature.txtRNA-level feature table (with ASE/hFDR/editing-related annotations). -
output_dir/all_feature.txtIntegrated feature table produced bymerge_feature. -
output_dir/all_feature.parquet
Mutation-prediction outputs¶
Typical outputs from mutation_prediction are:
output_dir/mutation_prediction/results/Sample_total_pred_truesites.vcfoutput_dir/mutation_prediction/results/Sample_total_pred_truesites_PASS.vcf
#CHROM POS ID REF ALT QUAL FILTER INFO
chr1 16562519 . C A . PASS DP=196;ALT_DP=24;VAF=0.122449;NUM_SPOTS=135;NUM_MUT_SPOTS=23;DNA_MUT_TYPE=A[C>A]A;RNA_MUT_TYPE=T[G>T]T
chr1 19682510 . G A . PASS DP=79;ALT_DP=17;VAF=0.21519;NUM_SPOTS=71;NUM_MUT_SPOTS=17;DNA_MUT_TYPE=T[C>T]T;RNA_MUT_TYPE=T[C>T]T
chr1 36393565 . G A . PASS DP=525;ALT_DP=89;VAF=0.169524;NUM_SPOTS=368;NUM_MUT_SPOTS=79;DNA_MUT_TYPE=T[C>T]C;RNA_MUT_TYPE=T[C>T]C
chr1 46675536 . G A . PASS DP=118;ALT_DP=20;VAF=0.160494;NUM_SPOTS=73;NUM_MUT_SPOTS=13;DNA_MUT_TYPE=G[C>T]T;RNA_MUT_TYPE=G[C>T]T
chr1 155234903 . C T . PASS DP=39;ALT_DP=3;VAF=0.0769231;NUM_SPOTS=39;NUM_MUT_SPOTS=3;DNA_MUT_TYPE=G[C>T]T;RNA_MUT_TYPE=A[G>A]CVisualization and validation¶
1) Lineage reconstruction with PhyloSOLID¶
After obtaining final predicted sites, you can run PhyloSOLID to infer lineage relationships.
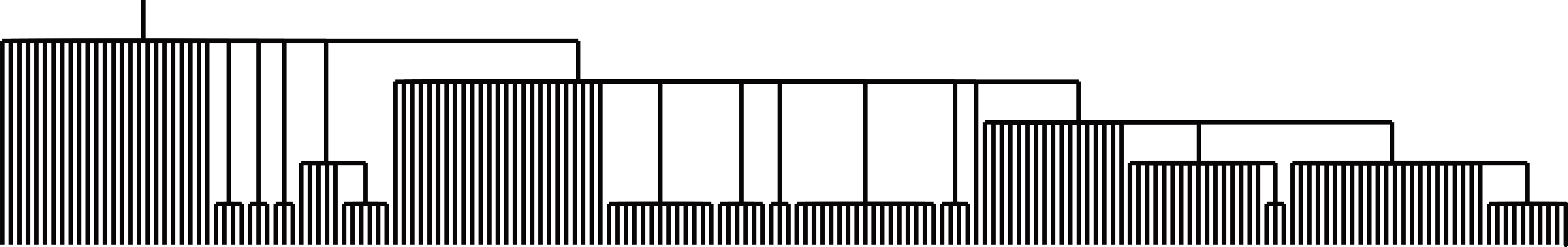
2) Spatial probability scatter plot¶
SpaceTracer supports spatial visualization of mutation probability and mutant/non-mutant spot patterns.
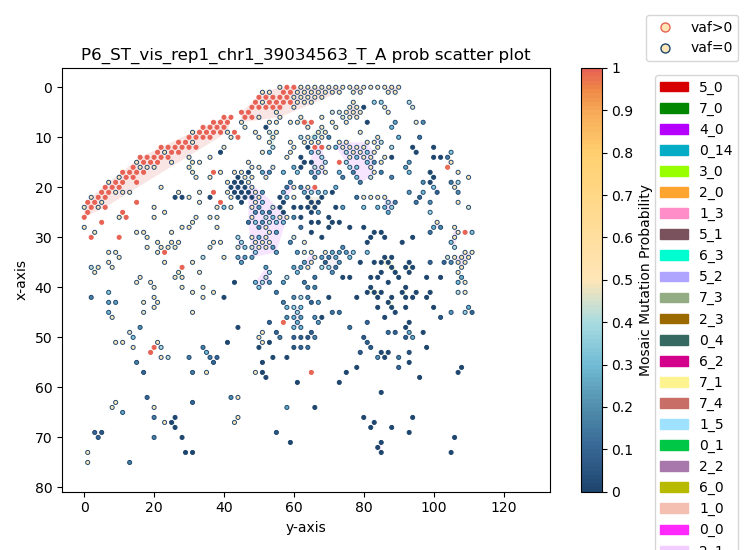
3) IGV validation views¶
To manually validate candidate calls, IGV snapshots can be used to compare mutant-like and non-mutant/reference-homozygous spots.
Mutant spots
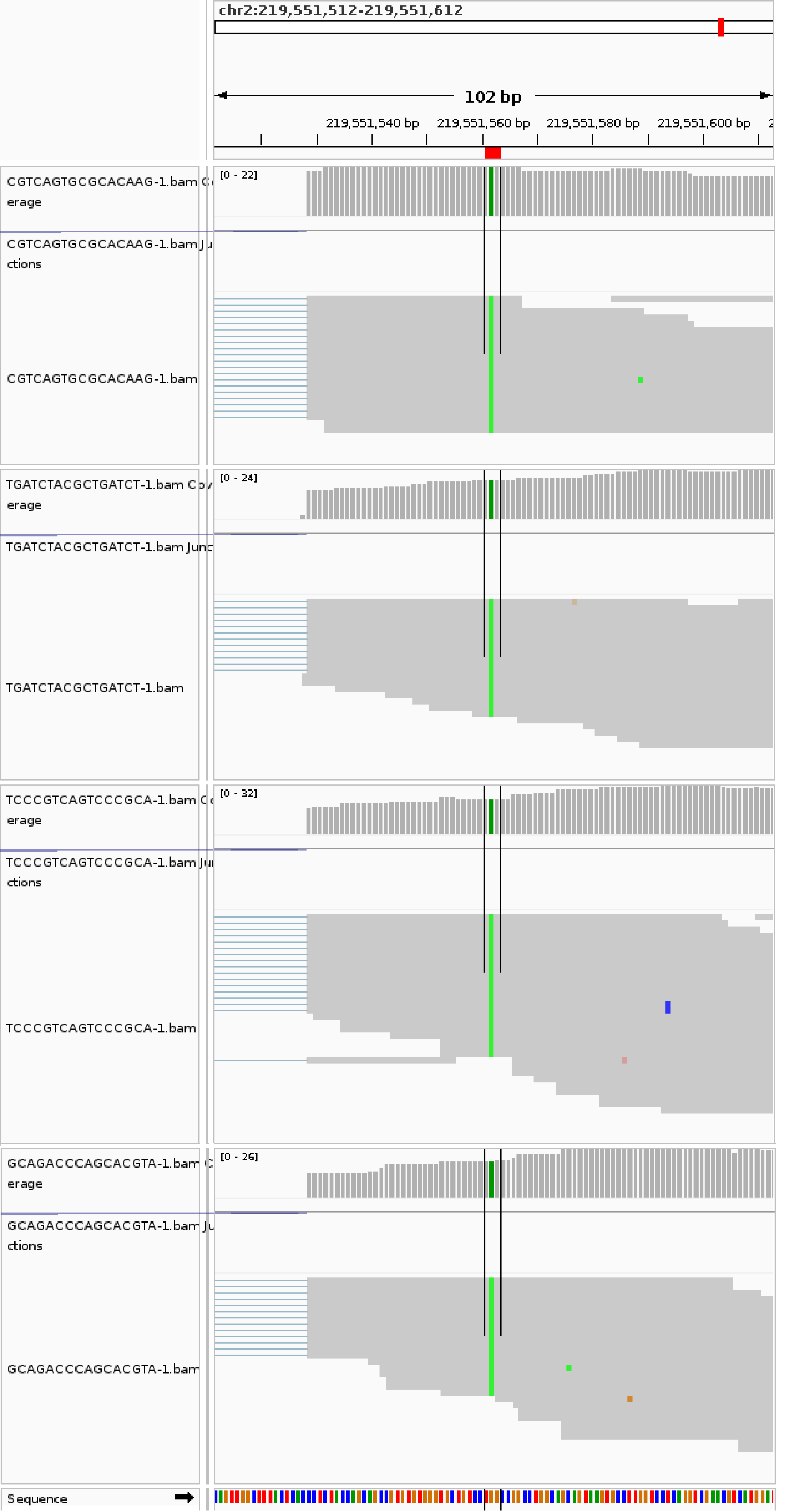
Non-mutant/reference-homozygous spots
